Single cell RNA-seq data analysis with Chipster
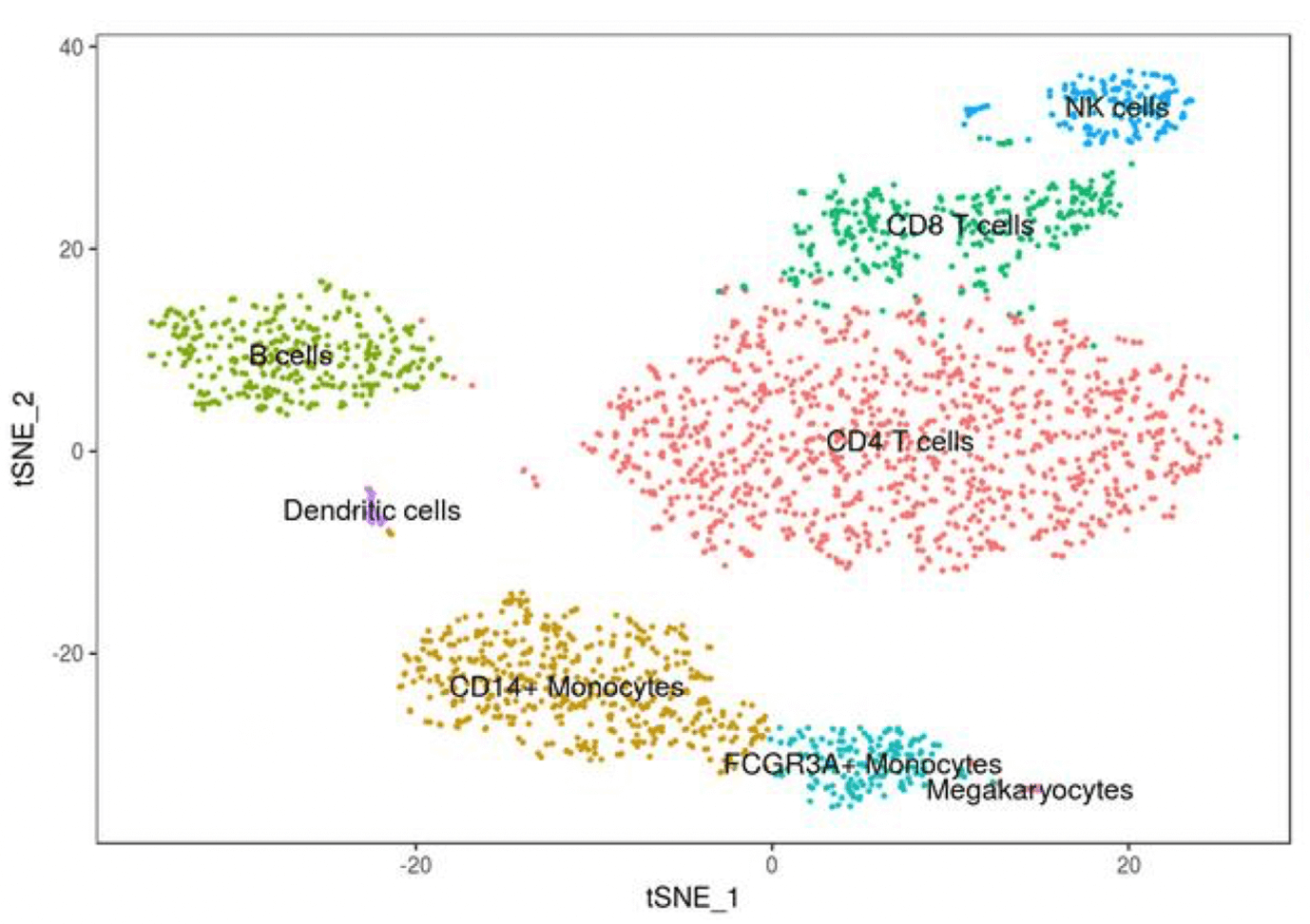
Description
This two half-days practical course (October 27, October 28) introduces single cell RNA-seq data analysis methods, tools and file formats to all researchers who are planning to use single cell RNA-seq data for their own projects. The course will cover the processing of transcript counts from quality control and filtering to dimensional reduction, clustering, and differential expression analysis. In addition, it will cover how to do integrated analysis of two samples.
Seurat v3 tools embedded in user-friendly Chipster software will be used for the practical tutorials and hands-on sessions.
Successful completion will be awarded with 0.5 ECTS.
Learning objectives
-
Obtaining an overview and discuss a variety of aspects of single cell RNA-seq data analysis
-
Gaining experience with Seurat v3 tools available in Chipster to undertake analysis of single cell RNA-seq data
-
Understanding the advantages and limitations of single cell RNA-seq data analysis
Requirements
The free and user-friendly Chipster software is used in the exercises, so no previous knowledge of Unix or R is required, and the course is thus suitable for everybody who is planning to use single cell RNA-seq.
Participants will be provided with the access to pre-recorded lessons two weeks before the online training as an introductionary overview of the main concepts and first steps with Chipster software. Participants will be expected to view these recordings and perform a short Chipster exercise before attending the online sessions.
Time Plan
Both days consist of interleaved lectures and hands-on exercises followed by a wrap-up and question session on each topic. A 15 min coffee break is held in between.
| Date | Time | Topic |
|---|---|---|
| Tue, 27 Oct | 8.45 | Virtual coffee & get together |
| 9.15 | Introduction to Chipster | |
| Introduction to single cell RNA-seq | ||
| Preprocessing and quality control | ||
| Dimensional reduction | ||
| 13.00 | End of day 1 | |
| Wed, 28 Oct | 8.45 | Virtual coffee & get together, re-cap |
| 9.15 | Clustering and finding marker genes, visualisation | |
| Analysis of two samples Integration Finding common cell types Finding conserved cluster marker genes Finding genes which are differentially expressed in a cell-type specific manner Visualisation |
||
| Conclusion & final discussion | ||
| 13.00 | End of course |
Registration
The registration is closed.
Speakers
Eija Korpelainen with Maria Lehtivaara
Address
The course will be held online. Access will be available for registered participants only.
